PLATFORm
AI allows us to make proteins that nature could if it had enough time.
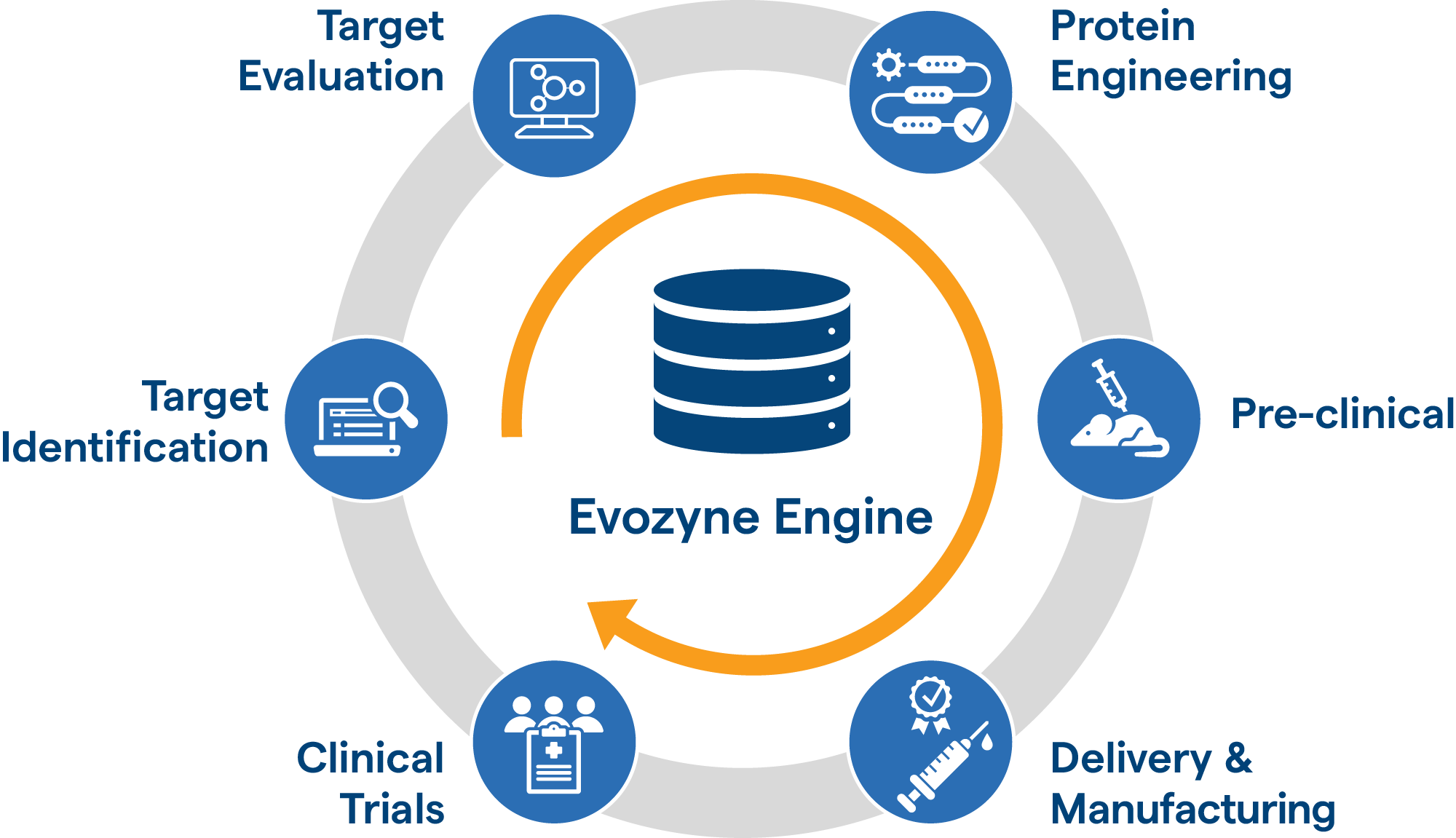
Evozyne uses the rules of evolution to design new molecules specifically to treat illnesses. By speeding up a natural process, we can potentially make breakthrough treatments for intractable diseases.
It’s a synergistic approach that encompasses our proprietary models, high-throughput assays and therapeutic area expertise. Leveraging these strengths means targets are generated in less than three months with lead candidates identified in under a year.
The insights gained during the discovery phase fill Evozyne’s proprietary data moat and result in a pipeline of potential new therapies.

